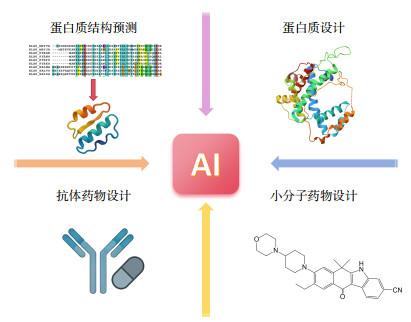
1 AI驱动蛋白质结构预测:AlphaFold3的突破与应用
1.1 AI在蛋白质结构预测中的应用
1.2 结构预测工具在药物研发中的最新研究成果
1.3 AlphaFold3的局限性与挑战
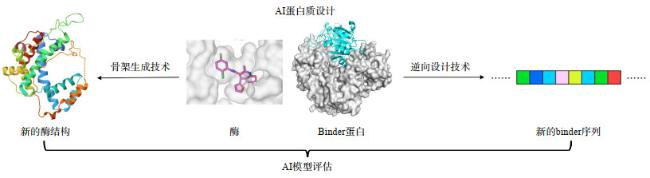
2 AI驱动蛋白质设计:从结构预测到功能创新
2.1 AI在蛋白质设计中的应用
2.2 AI驱动蛋白质设计的最新研究成果
2.3 AI驱动蛋白质设计面临的挑战
3 AI赋能抗体药物设计:加速研发进程,提升药物疗效
3.1 AI在抗体药物设计中的应用
表1 AI模型在抗体药物设计中的应用对比 |
| 模型类型 | 技术原理简述 | 主要应用环节 | 优势 |
| 扩散模型 | 通过逐步添加/去除噪声学习分布,实现蛋白三维结构的高精度生成 | 抗体骨架设计、CDR结构建模 | 可精确建模抗原表位几何约束,探索新结构构象 |
| 图神经网络 | 将分子结构建模为图,学习节点(残基)和边(相互作用)之间的拓扑关系 | 抗体−抗原互作预测、亲和力预测 | 能处理抗体−抗原结构复杂性,支持界面残基识别与突变筛选 |
| 生成对抗网络 | 编码抗体序列到潜在空间或训练对抗网络,生成结构合理的新抗体序列 | 抗体序列生成、亲和力多样性探索 | 可合成高质量且多样的抗体序列,辅助构建抗体库 |
| 语言模型 | 使用蛋白序列作为输入,基于自注意力机制学习序列和功能信息 | 序列优化、指导亲和力成熟 | 适用于大规模数据、快速评估突变影响 |
3.2 AI驱动抗体药物设计的最新研究成果
3.3 AI驱动抗体药物设计面临的挑战
4 AI助力小分子药物设计:加速发现、优化性质、提高效率
4.1 AI在小分子药物设计中的应用
4.2 AI驱动小分子药物设计的最新研究成果
4.3 AI驱动小分子药物设计面临的挑战
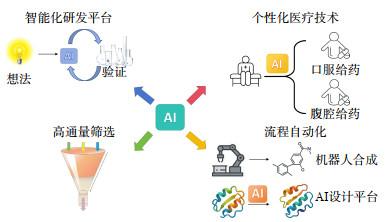
5 AI驱动药物研发的未来展望
表2 AI驱动药物研发进展与挑战总结 |
| 领域 | 关键进展 | 主要应用场景 | 现存挑战 | 未来发展方向 |
| 蛋白质结构预测 | AlphaFold3实现生物分子复合物结构预测 预测精度接近实验方法 突破性扩展至配体,核酸相互作用预测 | 靶点发现与验证 虚拟筛选与分子对接 抗体−抗原复合物解析 酶工程与药物重定位 | 动态行为预测能力不足 计算资源需求高 实验验证成本高 | 动态构象与相互作用预测 算法优化降低算力需求 多模态数据整合与平台开发 |
| 蛋白质设计 | RFdiffusion实现功能导向的从头设计 生成模型辅助结构改造 工业酶催化效率提升 | 新型治疗性蛋白质开发 酶从头设计 蛋白稳定性优化 合成生物学元件设计 | 非天然蛋白数据稀缺 功能预测准确性不足 实验验证成本高昂 | 多模态数据融合 动态行为预测模型开发 自动化设计−验证闭环平台 |
| 抗体药物设计 | RFantibody实现结构导向 抗体生成 AI亲和力优化 AI辅助可开发性预测 | 抗体人源化优化 亲和力成熟加速 双特异性抗体设计 稳定性与聚集倾向预测 | 免疫原性预测模型不足 动态表位预测困难 实验通量限制验证效率 | 多组学数据整合 可解释AI指导工程化改造 自动化高通量验证平台开发 |
| 小分子药物设计 | AI虚拟筛选与分子生成工具 AI化合物优化平台 ADMET预测模型 | 靶点识别与验证 全新分子骨架生成 合成路线智能规划 毒性预测 | 化学空间探索局限性 活性−选择性平衡困难 多靶点协同作用预测不足 | 知识图谱增强型生成模型 动态药效团建技术 自动化合成−测试闭环系统 |